

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.009G028200.1 |
| Family | CBM43 |
| Protein Properties | Length: 543 Molecular Weight: 60013.8 Isoelectric Point: 6.0529 |
| Chromosome | Chromosome/Scaffold: 09 Start: 2158922 End: 2162073 |
| Description | O-Glycosyl hydrolases family 17 protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM43 | 457 | 539 | 2e-31 |
| WCVATSQASRSNLQNALDWACGPGKADCSEIQPNQRCFQPDTLLAHASFAFNNYYRKNGATDASCSFRGNAIKVDRDPSYGNC | |||
| GH17 | 222 | 351 | 9.2e-32 |
| TRNTFDAQPDTVHFTLADTDLIISDIIAAETGWPTRGSRRPHASTHNAKTYNYSASLDSVDDYANIDNAQTYNTNLISHVMGGSGTPARPGANLDVYIFS LFNENLKQGPETERNFGLFYPDMTSVYNLV | |||
| CBM43 | 361 | 443 | 7.8e-32 |
| WCVASRQASNSALQNALDWACGPGKADCSALQPDQQCFEPNNLVSHASFAFNNYYQKNGLTDQACRFGGTGIVVYNDPSYGNC | |||
| GH17 | 2 | 218 | 0 |
| DADYLPSPEKVAQLVRNHNIQYLRIYNYRPEVLRAFSNTGIELMVGVPNADLSQFQSQPYVDSWLRTSILPYREATKITHITVGVEVTNSPDNAANLVVR AMRNVVSALKSANLQGKIKVSTPLSFGVLSKSYPPSEGAFNSGYEYVLRPLLDFLEENQSPFMVNLYPFYAIEDSSLDAVLFKSPSPIFVDQNTGLSYKN IFDAQLDAVFYAIANLN | |||
| Full Sequence |
|---|
| Protein Sequence Length: 543 Download |
| MDADYLPSPE KVAQLVRNHN IQYLRIYNYR PEVLRAFSNT GIELMVGVPN ADLSQFQSQP 60 YVDSWLRTSI LPYREATKIT HITVGVEVTN SPDNAANLVV RAMRNVVSAL KSANLQGKIK 120 VSTPLSFGVL SKSYPPSEGA FNSGYEYVLR PLLDFLEENQ SPFMVNLYPF YAIEDSSLDA 180 VLFKSPSPIF VDQNTGLSYK NIFDAQLDAV FYAIANLNFR TTRNTFDAQP DTVHFTLADT 240 DLIISDIIAA ETGWPTRGSR RPHASTHNAK TYNYSASLDS VDDYANIDNA QTYNTNLISH 300 VMGGSGTPAR PGANLDVYIF SLFNENLKQG PETERNFGLF YPDMTSVYNL VFPGKGTGRS 360 WCVASRQASN SALQNALDWA CGPGKADCSA LQPDQQCFEP NNLVSHASFA FNNYYQKNGL 420 TDQACRFGGT GIVVYNDPSY GNCIYNVKSR DPNGRTWCVA TSQASRSNLQ NALDWACGPG 480 KADCSEIQPN QRCFQPDTLL AHASFAFNNY YRKNGATDAS CSFRGNAIKV DRDPSYGNCI 540 YH* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam07983 | X8 | 3.0e-18 | 456 | 528 | 82 | + X8 domain. The X8 domain domain contains at least 6 conserved cysteine residues that presumably form three disulphide bridges. The domain is found in an Olive pollen allergen as well as at the C-terminus of several families of glycosyl hydrolases. This domain may be involved in carbohydrate binding. This domain is characteristic of GPI-anchored domains. | ||
| pfam07983 | X8 | 2.0e-18 | 360 | 432 | 78 | + X8 domain. The X8 domain domain contains at least 6 conserved cysteine residues that presumably form three disulphide bridges. The domain is found in an Olive pollen allergen as well as at the C-terminus of several families of glycosyl hydrolases. This domain may be involved in carbohydrate binding. This domain is characteristic of GPI-anchored domains. | ||
| smart00768 | X8 | 2.0e-34 | 456 | 541 | 86 | + Possibly involved in carbohydrate binding. The X8 domain, which may be involved in carbohydrate binding, is found in an Olive pollen antigen as well as at the C terminus of family 17 glycosyl hydrolases. It contains 6 conserved cysteine residues which presumably form three disulfide bridges. | ||
| smart00768 | X8 | 3.0e-36 | 360 | 445 | 86 | + Possibly involved in carbohydrate binding. The X8 domain, which may be involved in carbohydrate binding, is found in an Olive pollen antigen as well as at the C terminus of family 17 glycosyl hydrolases. It contains 6 conserved cysteine residues which presumably form three disulfide bridges. | ||
| pfam00332 | Glyco_hydro_17 | 5.0e-80 | 1 | 352 | 357 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002283473.1 | 0 | 2 | 445 | 33 | 449 | PREDICTED: similar to beta-1,3-glucanase [Vitis vinifera] |
| RefSeq | XP_002283473.1 | 2e-33 | 456 | 541 | 364 | 449 | PREDICTED: similar to beta-1,3-glucanase [Vitis vinifera] |
| RefSeq | XP_002330523.1 | 0 | 2 | 447 | 34 | 453 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002514201.1 | 0 | 2 | 445 | 33 | 451 | Glucan endo-1,3-beta-glucosidase precursor, putative [Ricinus communis] |
| RefSeq | XP_002514201.1 | 5e-34 | 456 | 541 | 366 | 451 | Glucan endo-1,3-beta-glucosidase precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 1 | 352 | 7 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_D | 0 | 1 | 352 | 8 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_C | 0 | 1 | 352 | 8 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_B | 0 | 1 | 352 | 8 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_A | 0 | 1 | 352 | 8 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
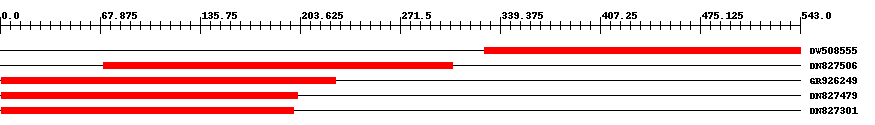 | ||||
| Hit | Length | Start | End | EValue |
| DW508555 | 215 | 329 | 543 | 0 |
| DN827506 | 238 | 70 | 307 | 0 |
| GR926249 | 228 | 1 | 228 | 0 |
| DN827479 | 202 | 1 | 202 | 0 |
| DN827301 | 199 | 1 | 199 | 0 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
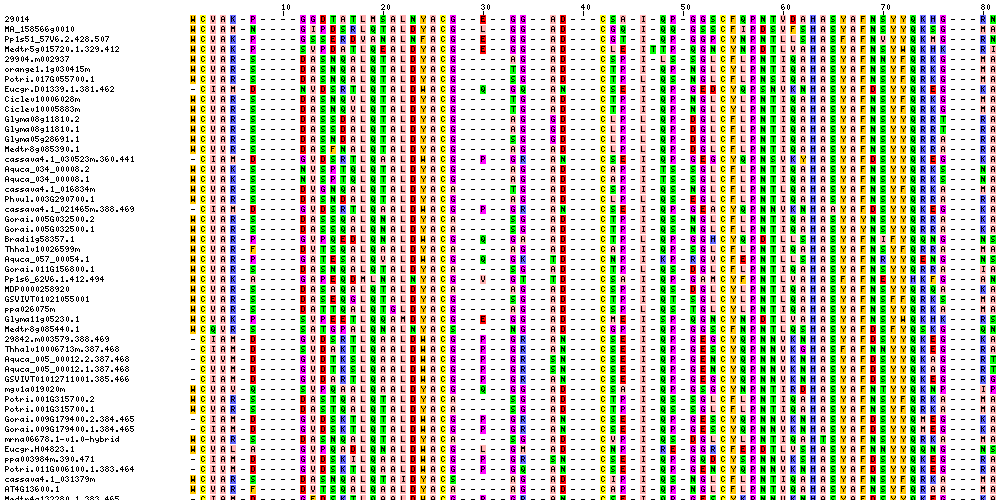 |